Speculation about translocation mechanism of carbon nanotubes into cells
Cell membranes represent a serious protective barrier for external molecules, proteins, nanoparticles and drugs. This barrier is quite efficient in protecting the interior of the cells. However, large nanoscale objects, single-walled carbon nanotubes (SWNTs) have been found inside the cells both in direct and indirect biological experiments. Such experiments suggest that carbon nanotubes can efficiently penetrate
into the cells but very little can be said about the pathway and the entry
mechanism regulating their internalization.
Do we believe in the spontaneous translocation through the piercing mechanism envisaged by many researchers?
Cartoon view of guessed interaction of carbon nanotubes with cells:
In order to check the validity of the spontaneous translocation through the bilayer and get some insight into this mechanism, we have performed series of SCMF calculations for different positions of perpendicularly oriented SWNT with respect to the DMPC phospholipid bilayer plane for different diameters of the SWNT and interaction parameters between the SWNT and the core of the bilayer.
This method gives the equilibrium free energy of the insertion of the nanotube into bilayer, concentration profiles of the heads and tails in the bilayer, the concentration profiles of the phospholipid bilayer which can demonstrate the structural rearrangements of phospholipids induced by the SWNT at the molecular level and the precise measurements of the equilibrium free energy change for each position of the SWNT. In addition, the SCMF theory gives the probabilities of each phospholipid conformation in the corresponding mean fields.
Carbon nanotubes, by their chemical structure, can be considered as hydrophobic cylinders. Such objects in aqueous solutions tend to aggregate in bundles, which makes it difficult to observe individual SWNT in solution. In order to prevent such aggregation, the surface of the SWNT is often treated with acids or functionalized in such a way that they become slightly hydrophilic. That is why, the energy per contact of the hydrophobic bead with the nanotube, in our calculations changes from 0, representing steric repulsion, to 6.3 kT, which corresponds to strong attraction.
The method allows for visualization of the mean field solution by means of a mean field "snapshot" (the set of most probable configurations).
and the equilibrium concentration profiles.
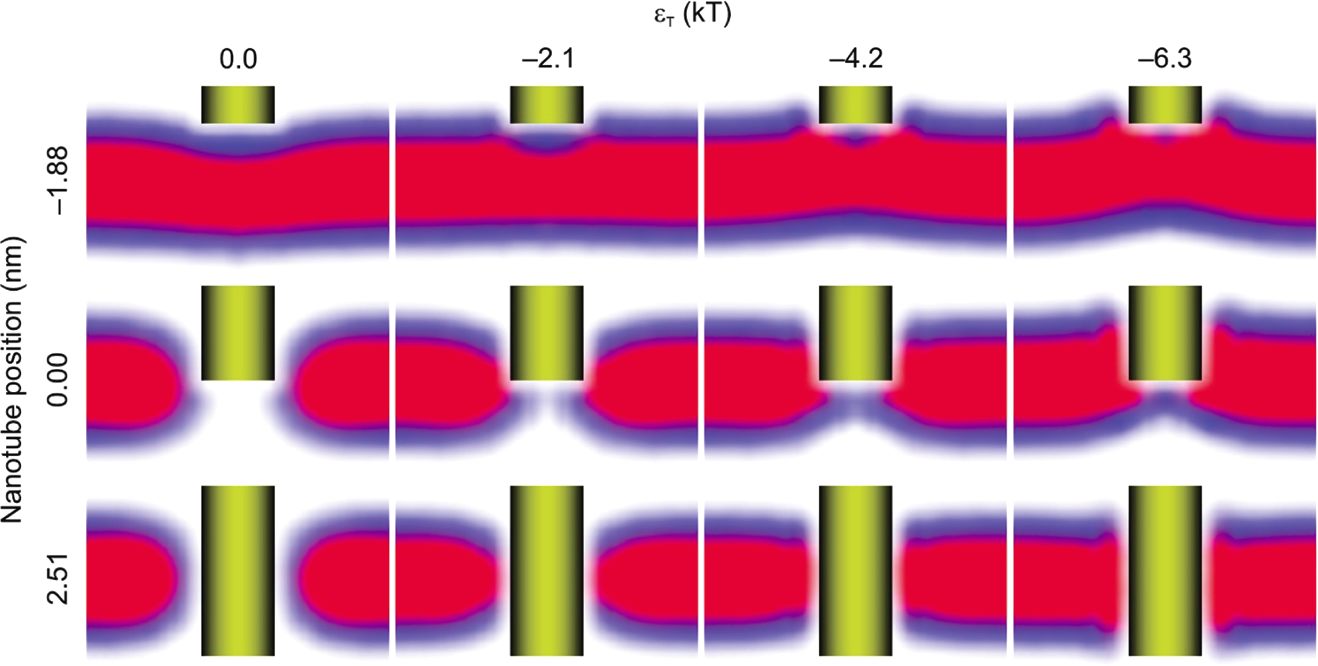
The result of calculations is shown for a SWNT of 1 nm diameter. Here the SWNT position -3.14 nm corresponds to the SWNT at the bilayer surface, while the position 3.14 nm corresponds to fully inserted SWNT.
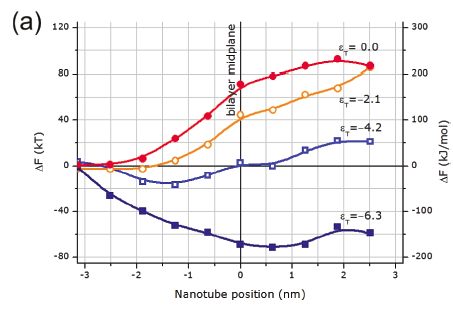
Our results suggest that hydrophilic or noninteracting SWNTs would require an energy of order of hundreds of kT to cross the phospholipid bilayer in perpendicular orientation. Furthermore, perpendicular orientation is less disruptive for the bilayer, and thus, other orientations would require even higher energies. In turn, more hydrophobic, interacting SWNTs are attracted by the hydrophobic cores, what hinders their translocation through the bilayers and renders difficult the separation of the nanotube from the bilayer simply by thermal motion.
Thus, our results for model phospholipid bilayers may suggest that experimentally observed translocation of SWNTs into cells is probably due to other energy-dependent translocation mechanisms, such as, for example, endocytosis.
Another conclusion is that in living cells the cytoplasm contains biomolecules (proteins and DNA) that can spontaneously adsorb onto nanotube, thus modifying the surface properties as shown in this example:
Full version:
Sergey Pogodin and Vladimir A. Baulin, "Can a Carbon Nanotube Pierce through a Phospholipid Bilayer?", ACS Nano, 2010, 4 (9), pp 5293–5300
Sergey Pogodin, Nigel Slater and Vladimir A. Baulin, "Surface Patterning of Carbon Nanotubes Can Enhance Their Penetration through a Phospholipid Bilayer", ACS Nano, 2011, 5 (2), pp 1141–1146

